Magna cum Laude Merit Award for research to detect the progress of diseases such as multiple sclerosis
The researchers’ imaging technique is fast, accurate, and reproducible
An interdisciplinary research team at the University of Michigan has developed a faster way to track the progress of neurodegenerative diseases, such as multiple sclerosis. Their work has been recognized both for its improved methodology, as well as its reproducibility, making the work more readily available to researchers around the world.
The team, led by Jeff Fessler, the William L. Root Collegiate Professor of EECS, aims to dramatically reduce the time it takes to take a scan of the brain’s myelin sheath, which is an important indicator of the brain’s neurological health. Myelin sheaths surround and protect nerve cells. The relative health of myelin sheaths can be determined by estimating the myelin water fraction (MWF).
“The standard method for MWF imaging is a multi-echo spin-echo (MESE) technique that is just too slow to be used routinely in the clinic,” said Fessler in an interview with teammates Steven Whitaker, a doctoral student in electrical and computer engineering, and Dr. Jon-Fredrik Nielsen, associate research scientist in the Department of Biomedical Engineering.
An alternative method called small-tip fast recovery (STFR) had been developed by the team to speed the time it takes to scan the brain, but the results were less accurate because they did not consider the exchange of myelin and non-myelin water in the method.
 Enlarge
Enlarge
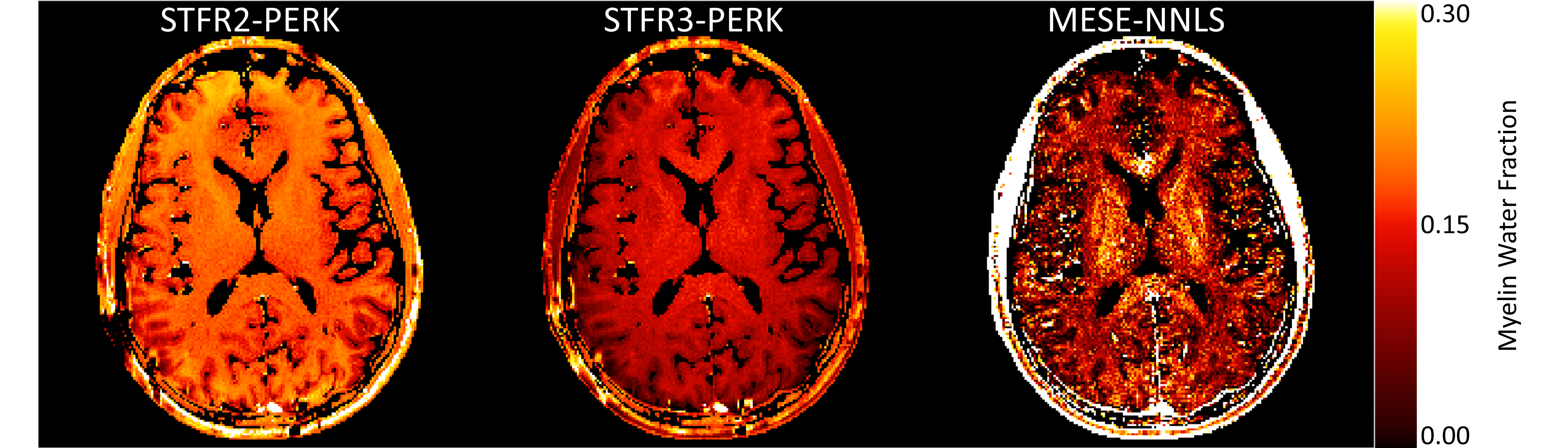
In the new method, the Michigan team used a three-compartment tissue model consisting of myelin water, non-myelin water, and a macromolecular pool. Their results came much closer to the MESE-based estimates, while reducing the scan time from 36 min 11 sec to 7 min 14 sec.
In addition to publishing their results, the team documented their methodology and shared the complete source code and data in a way that makes it simple for other researchers to duplicate their work.
“I’ve been benefiting from open source tools for my entire career, and I feel that this is a way of giving something back,” said Fessler in the same interview. This is not new for Fessler, who regularly posts his algorithms on the Internet.
The team used Nielsen’s framework for rapid prototyping of MR pulse sequences in the research. Called TOPPE, it is also openly available, and used by many MRI researchers at U-M and beyond.
The research, “Myelin Water Imaging Using STFR with Exchange” by Steven T. Whitaker, Gopal Nataraj, Jon-Frederik Nielsen, and Jeffrey A. Fessler, was presented at the 2020 International Society for Magnetic Resonance in Medicine (ISMRM) & Society for MR Radiographers & Technologists (SMRT) Virtual Conference & Exhibition. Whitaker received the Magna cum Laude Merit Award at the conference for his presentation of the research.
The journal Magnetic Resonance in Medicine featured the research, published earlier in the year, in August 2020 with two interviews:
– Reproducible Research Insights with Steven Whitaker, Jon-Fredrik Nielsen and Jeff Fessler (August 28, 2020)
– Q&A with Steven Whitaker, Jon-Fredrik Nielsen and Jeff Fessler (August 21, 2020)
The fourth author, Dr. Gopal Nataraj, recently graduated from Michigan and is a postdoctoral research fellow at Memorial Sloan Kettering Cancer Center.
 MENU
MENU 
